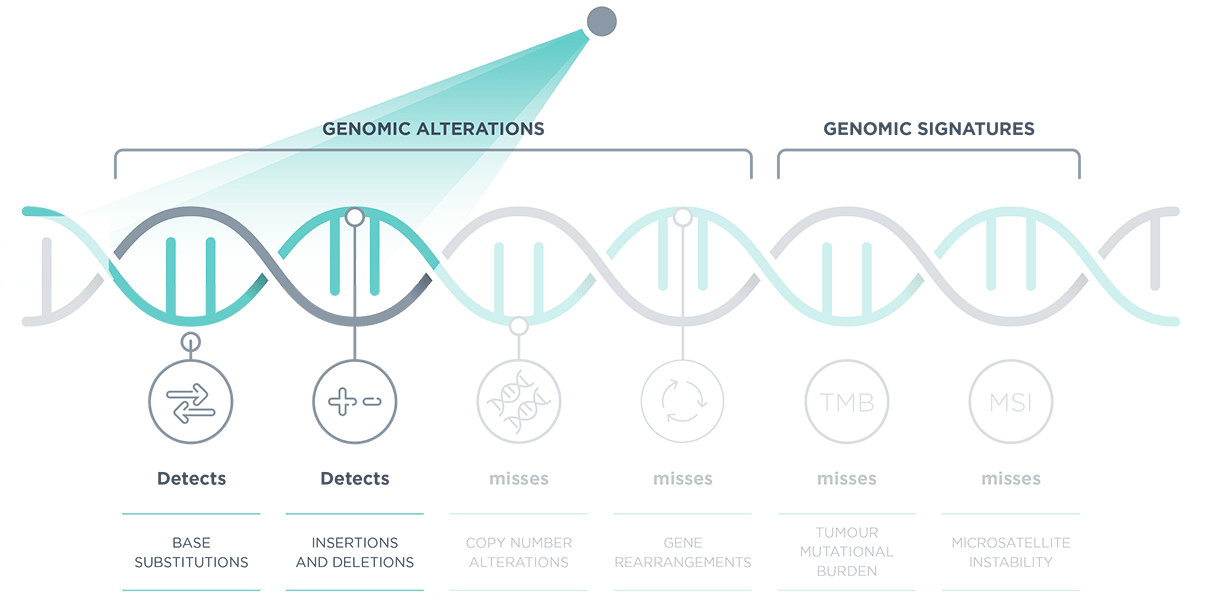
*Comprehensive genomic profiling provides prognostic, diagnostic and predictive insights that inform treatment decisions for individual patients across all cancer types.
†TMB reported by FoundationOne CDx and FoundationOne Heme. bTMB reported by FoundationOne Liquid CDx. MSI reported by FoundationOne CDx and FoundationOne Heme, MSI-H reported by FoundationOne Liquid CDx.
‡Therapies contained in the EU version o f the report may have been approved through a centralised EU procedure or a national procedure in an EU Member State.
§For additional information on the NCCN categories please refer to the NCCN Compendium® (www.nccn.org).
¥Clinical validation demonstrated concordance with the following companion diagnostics: cobas® EGFR Mutation Test. Ventana ALK (DSF3) CDx Assay, Vysis ALK BreakApart FISH Probe Kit, therascreen® KRAS RGQ PCR Kit, Dako HER2 FISH PharmDx® Kit, cobas® BRAF V600 Mutation Test, THxlD® BRAF kit. For more information, please see the FoundationOne®CDx Technical Specifications available at: www.rochefoundationmedicine.com/f1cdxtech.
¶Clinical validation demonstrated concordance with the following diagnostics: cobas® EGFR Mutation Test v2, a tumor tissue polymerase chain reaction-based clinical trial assay (CTA), and an externally validated circulating cell-free DNA-based next-generation sequencing assay. For more information please see the Found ationOne Liquid®CDx technical Specifications available at: www.eifu.online/FMl/190070862.
bTMB, blood Mutational Tumour Burden; FDA, US Food and Drug Administration; MSI, Microsatellite Instability; NCCN, Nat ional Comprehensive Cancer Network; NGS, next generation sequencing; TMB, Tumour Mutational Burden.
- FoundationOne®CDx Technical Specifications, 2018. Available at: www.rochefoundationmedicine.com/f1cdxtech (Accessed August 2020).
- Data on file: FoundationOne Liquid CDx Technical Specifications, 2020. Available at: http://www.eifu.online/FMI/190070862 (Accessed August 2020).
- FoundationOne®Heme Technical Specifications, 2017. Available at: www.foundationmedicine.com/genomic-testing/foundation-one-heme (Accessed August 2020).
- Frampton GM et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol ; 31: 1023–1031. 2013.
- Data on file: Clinical and analytical validation data file for FoundationOne Liquid CDx.
- Clark TA et al. Analytical Validation of a Hybrid Capture-Based Next-Generation Sequencing Clinical Assay for Genomic Profiling of Cell-Free Circulating Tumor DNA. J Mol Diagn ; 20: 686–702. 2018
- He J et al. Integrated genomic DNA/RNA profiling of hematologic malignancies in the clinical setting. Blood; 127: 3004–3014. 2016
- Suh JH et al. Comprehensive Genomic Profiling Facilitates Implementation of the National Comprehensive Cancer Network Guidelines for Lung Cancer Biomarker Testing and Identifies Patients Who May Benefit From Enrollment in Mechanism-Driven Clinical Trials. Oncologist ; 21: 684–691. 2016
- Chalmers ZR et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med ; 9: 34. 2017
- Rozenblum AB et al. Clinical Impact of Hybrid Capture-Based Next-Generation Sequencing on Changes in Treatment Decisions in Lung Cancer. J Thorac Oncol; 12: 258–268. 2017
- Schrock AB et al. Comprehensive Genomic Profiling Identifies Frequent Drug-Sensitive EGFR Exon 19 Deletions in NSCLC not Identified by Prior Molecular Testing. Clin Cancer Res; 22: 3281–3285. 2016
- Ross JS et al. Comprehensive genomic profiling of epithelial ovarian cancer by next generation sequencing-based diagnostic assay reveals new routes to targeted therapies. Gynecol Oncol ; 130: 554–559. 2013
- Hall MJ et al. Evaluation of microsatellite instability (MSI) status in 11,573 diverse solid tumors using comprehensive genomic profiling (CGP). J Clin Oncol ; 34: 1523–1523. 2016
- FoundationOne®CDx Sample Report. Available at: https://www.foundationmedicine.nl/content/dam/rfm/sample-reports/f1cdx/eu_version_-_ema_without_page_1/F1CDx%20EU%20Sample%20Report%20(Lung).pdf (Accessed August 2020).
- A search for "Foundation Medicine"[Affiliation] on the NCBI database resulted in 518 publications, as of April 2020. Available at: https://www.ncbi.nlm.nih.gov/pubmed/?term=%22Foundation+Medicine%22%5BAffiliation%5D (Accessed August 2020).
- Foundation Medicine Publications. Available at: https://www.foundationmedicine.com/publications (Accessed August 2020).
- FoundationOne®CDx FDA Approval, 2017. Available at:
https://www.accessdata.fda.gov/cdrh_docs/pdf17/P170019a.pdf (Accessed August 2020). - FoundationOne®CDx Clinical Validation, 2017. Available at: http://www.foundationmedicine.com/genomic-testing/foundation-one-cdx (Accessed August 2020).
- FoundationOne Liquid CDx FDA Approval, 2020. Available at: https://www.foundationmedicine.com/press-releases/445c1f9e-6cbb-488b-84ad-5f133612b721 (Accessed August 2020).